Meet Whitman students, past and present, who have worked in the lab [click here!]
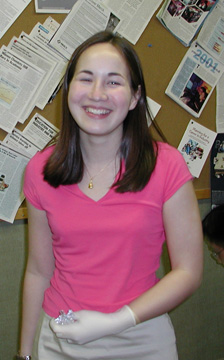

Dan Vernon Homepage - Daniel M. Vernon laboratory, Whitman College
The Vernon Lab
Research in plant functional genomics and molecular biology at Whitman College
Research:
Scroll down for info on our research interests and current and past projects
Publications & Presentations
C.V.s - Daniel M. Vernon (.pdf); Nancy R. Forsthoefel
Teaching:
ASPB 2009 Education Presentation:
An Adaptable Undergraduate Molecular Biology Lab Module that Integrates Use of Genomic Resources with Bench Experiments
Team Weed:
Meet Whitman students, past and present, who have worked in the lab [click here!]
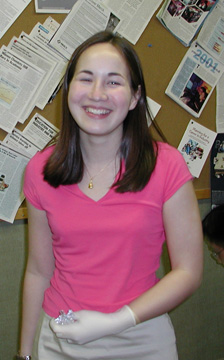
![]()
![]()

OUR RESEARCH: |
||
Our lab identifies plant genes and investigates their functions. We are interested in genes involved in reproduction and development (the process by which a complex, multicellular plant develops from a single cell). We are also interested in the molecular evolution of the genes we study. We are grateful for support from the National Science Foundation [www.nsf.gov] and for previous support from the USDA and W.J. Murdock Charitable Trust. We do our research on a small weed, Arabidopsis thaliana, a member of the mustard family popular for research on plant genetics, genomics, and development. Like the human genome, the Arabidopsis genome has been the focus of a massive genome sequencing project, and its DNA has been completely sequenced, just like that of the human genome. Arabidopsis has therefore become one of a small, select group of model species that biologists focus on to identify genes and learn about genomes. Determining the functions all of this plant's genes, and figuring out how they work together to build and operate the plant, is now a major goal of plant biologists. Our work follows up on the Arabidopsis genome project by defining related groups of genes, determining their boundaries, surveying their expression, and investigating their functions. More detailed information on Arabidopsis and its genome is available at The Arabidopsis Information Resource (TAIR; http://www.arabidopsis.org/) |
  Arabidopsis thaliana: A model system for studies of plant biology. You gotta love it! |
|
|
As one strategy to figure out what different genes do, we study gene knock-out mutants. These are mutants in which a specific gene has been disrupted by insertion of a foreign piece of DNA. One can learn where a gene is active and what processes it is important for by observing what happens when it is screwed up. We are working on mutants for two different kinds of genes. The first encodes Plant Intracellular Ras-group-related Leucine-rich repeat proteins (PIRLs), which are a familyof Leucine-Rich Repeat proteins found only in plants, but which are related to animal LRR proteins involved in gene regulation and cell signaling. The second group of genes we study are Pentatricopeptide Repeat (PPR) genes, many of which encode proteins that act in post-transcriptional steps of gene expression in organelles (plastids and mitochondria). We work on a sub-set of PPR genes that are essential for embryogenesis. Much of our past research was also on plant embryo development. Scroll down for sample data nuggets from these projects and others. Information on the PIRLs is available in these publications from our lab: Forsthoefel et al, 2005; Vernon & Forsthoefel 2002; Forsthoefel et al. 2010 (submitted); and several meeting presentations [e.g., Forsthoefel et al, 2006; Forsthoefel et al, 2009]. Our work with PPR genes is described in these papers and abstracts: Cushing et al, 2005; Vernon et al, 2008; and Anderson et al, 2004; . Our lab is at Whitman College, and Whitman undergraduate students make important contributions to the research. The Publication, C.V., and Team Weed links here (or above) lead to more information on our research and members of the lab. Some representative examples of research findings & topics are shown below, with links to corresponding publications. |
|
|
|
Graphs comparing intron numbers for PPR genes (on left) to a set of control genes of similar size (right). Most PPR genes have no introns. This is part of our ongoing study of PPR gene evolution. PPR genes are found in all eukaryotes (plants, animals, fungi, etc) but in plants they are one of the biggest classes of genes. (Arabidopsis has >400 PPR-encoding genes). How did this family expand so much in plants? Our hypothesis is that early in plant evolution, PPRs multiplied via RNA intermediates: they got transcribed to RNA, then reverse-transcribed back into DNA and re-inserted into the genome. One hallmark of this type of gene duplication is genes that lack introns, since introns are removed from RNA just after transcription. [Anderson et al, 2004] |
|
|
 Regulation of embryo morphology and organ patterning Regulation of embryo morphology and organ patterning The above photographs show examples of twin1 mutant seedlings with abnormal cotyledon patterning and morphology. Panel 1 shows a wild-type Arabidopsis seedling, with 2 identical cotyledons (specialized leaves that form during embryogenesis). Panels 2-6 show various twin1 mutant seedlings with single, fused, or triple cotyledons. These results showed that the TWIN1 gene is important for proper organ patterning at the embryonic shoot apex, as well its previously defined function in the developing embryo. (From Vernon et al., 2001) |